- Home
- News
- Spotlight on Science
- How cohesin ensures...
How cohesin ensures sister chromatids segregate correctly
15-01-2019
In cell division, after replication of the cell’s chromosomes, the two copies, called sister chromatids, must be kept together to ensure that each daughter cell receives an equal complement of chromosomes. The protein complex cohesin keeps the sister chromatids together, but how it interacts with the DNA was unknown. A crystal structure of a part of the yeast cohesin complex bound to DNA has now been solved and reveals how the complex engages the DNA double helix providing insights into a fundamental cell process important in many diseases.
An essential part of cell division is the duplication of the genetic material. In higher organisms, DNA is packaged into chromosomes. Before the cell physically divides, the newly copied chromosome is kept tightly bound to its sister in order to make sure that when the cell separates, each half has one copy of each chromosome. A protein complex, called cohesin, that is conserved from single celled organisms all the way to humans, is responsible for keeping the sister chromosomes together by forming a ring around the DNA. Despite the importance of this complex, exactly how it interacts with the DNA in the chromosomes was unknown.
A structure of a part of the cohesin complex bound to double stranded DNA has now been revealed. The regions of the cohesion complex responsible for binding DNA were determined in the yeast complex and the components called Scc3 and Scc1 were purified. The mechanism of DNA binding was elucidated by determining the structure of this complex with double stranded DNA.
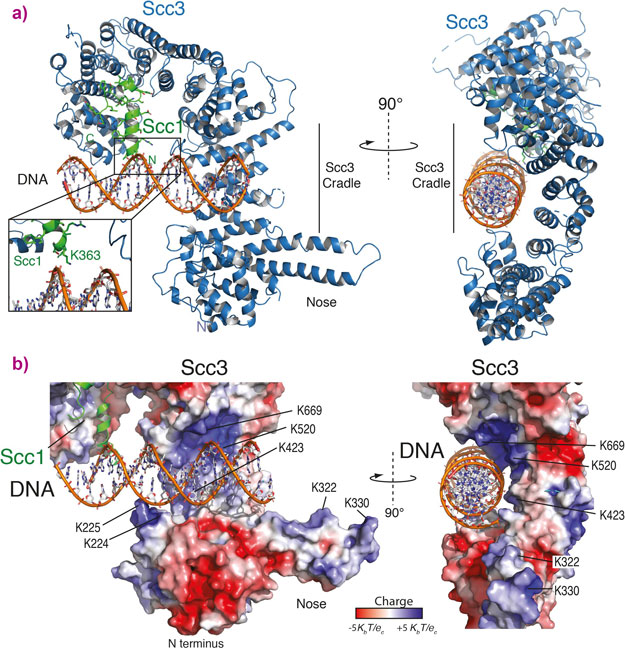
The structure shows a cradle of positively charged residues that are able to bind DNA (Figure 1). The positively charged residues in the Scc3/Scc1 complex directly interact with the negatively charged backbone of the DNA double helix showing that it binds double stranded DNA, but is not sequence specific. Further experiments were then carried out in yeast cells showing that cohesin can no longer load onto DNA if the DNA binding cradle is disrupted.
The Scc3/Scc1 complex is large and flexible and it proved very difficult to obtain well-diffracting crystals. As well as extensive screening of protein constructs, DNA fragments of different lengths and overhangs were screened. The fully-automated beamline MASSIF-1 (ID30A-1) was the perfect tool for screening a large number of crystals. When screening new DNA fragments, only a few crystals in every hundred would diffract sufficiently to provide molecular details needed to define the protein DNA interaction. The crystals also diffracted anisotropically, with the best direction extending to 3.6 Å and only ~5.7 Å in the worst direction. This meant that the sequence of the protein had to be defined by scanning mutations containing seleno-methionine in order to use the anomalous signal to define the register of the protein sequence. This also involved considerable screening to collect the best data for each mutant. In the end, over 800 crystals were required to produce the final model of the protein/DNA complex. The ability of the beamline to automatically screen crystals without any human involvement was essential in allowing the team to concentrate on quickly refining crystallisation conditions and selecting the best data sets. However, an important quality of the beamline was also the optimisation of data collection. In this case, the beam diameter was dynamically changed to match the volume of the crystal and X-ray doses optimised to the diffraction limit and volume of each selected crystal [1]. This ensured that the best possible data were collected for each crystal.
Principal publication and authors
Structural basis for Scc3-dependent cohesin recruitment to chromatin, Y. Li (a), K. Muir (a), M.W. Bowler (a), J. Metz (b), C. Haering (b) & D. Panne (a,c), eLife 7, e38356 (2018); doi: 10.7554/eLife.38356.
(a) EMBL, Grenoble Outstation (France)
(b) Cell Biology and Biophysics Unit, EMBL, Heidelberg (Germany)
(c) Leicester Institute of Structural and Chemical Biology, Department of Molecular and Cell Biology, University of Leicester (UK)
References
[1] O. Svensson, M. Gilski, D. Nurizzo & M.W. Bowler, Multi-position data collection and dynamic beam sizing: recent improvements to the automatic data-collection algorithms on MASSIF-1, Acta Cryst. D74, 433-440 (2018); doi: 10.1107/S2059798318003728.
Top image: During cell division, cohesin directly engages DNA (purple) and holds the replicated chromosome together. IMAGE: Nasser Rusan/National Institutes of Health (CC BY 2.0).